- Blog
- Ggn Wiz Khalifa
- Wintergreen Beard Oil Recipe
- Ptgui Pro 11.6 Crack
- Muvizu Play Plus Free Download
- Got S08e03 Kickass
- Smeg Scp108r-8 Price
- Ebook Oracle 11g English Pdf
- Corvus Games Terrain
- Dragon ball z dublado episodeo 65
- Command and conquer generals zero hour mods
- Mukis kitchen cartoon
- Logic pro 8 updates
- Radious total war mod need units
- Dmca nintendo 500 games
- Login fifa web app
- Is there a audio recorder on mac macbook pro
- Demon king daimao episode 9 english dub
- You crack me up meme
- Paul tekken 3
- Now 4 cd
- Read free anne mather books online
- Hum saath saath hain movie tabu marrige
- The walking dead a new frontier telltale
- Thomas trainz music video
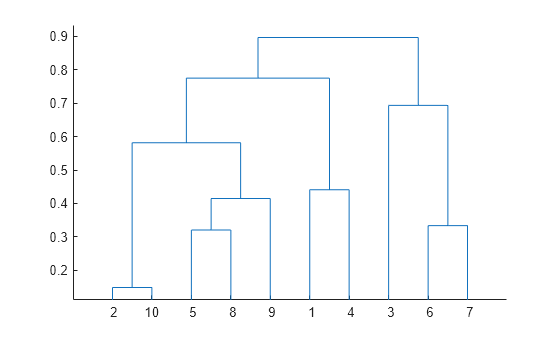
- Dendrogram construction bioedit
- Arabic pack for mathtype 6-9
- Argentine remington rolling block rifle
- Taylormade m2 driver
- Mendeley plugin word
- Pokemon sun and moon booster box
- Refx nexus 2-6-5 working activation key
- Asap rocky forever copy trailer
- Imagine dragons album night visions
- The hate you give soundtrack
- Planet coaster tutorial
- Office 2016 professional plus

The diluted cultures and real drinking water samples were tested by the microarray with 100% accuracy. The results were specific and reproducible, with a detection sensitivity of 0.1 ng DNA or 10 4 CFU/ml achieved for pure cultures of each target organism. Testing was carried out against a total of 218 bacterial strains, including 53 representative strains, 103 clinical isolates and 62 strains of other bacterial species belonging to 10 genera and 48 species.
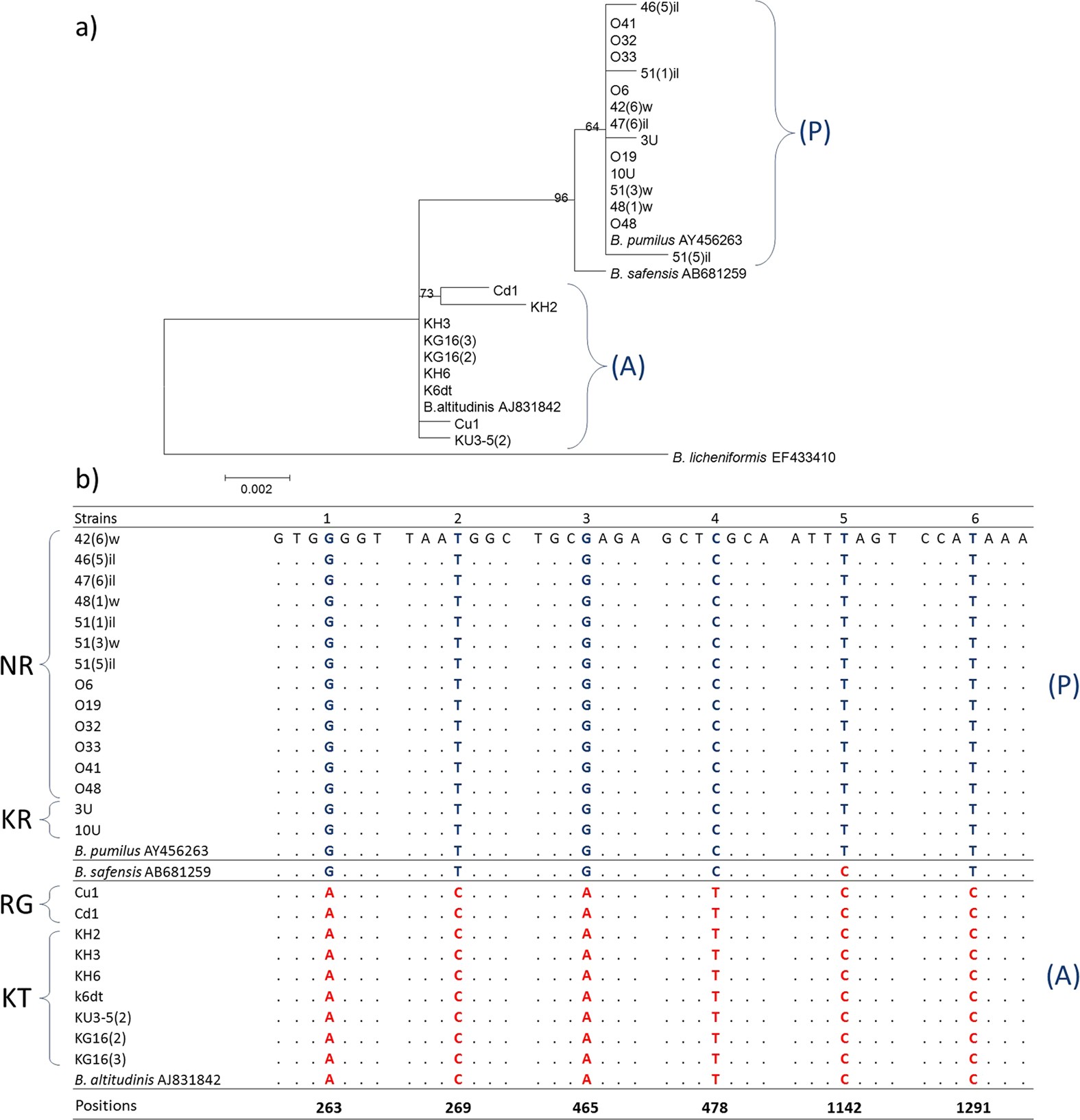
This new diagnostic contains 26 specific probes and can simultaneously detect Aeromonas hydrophila, Klebsiella pneumoniae, Legionella pneumophila, Pseudomonas aeruginosa, Salmonella spp., Shigella spp., Staphylococcus aureus, Vibrio choleraeo, Vibrio parahaemolyticus, Yersinia enterocolitica and Leptospira interrogans. Published online ahead of print on as DOI 10.1099/ ijs.0.63722-0. Evolutionary distances were calculated using the Jukes & Cantor evolutionary model and a phylogenetic tree was Abbreviation: rep-PCR, repetitive-element polymerase chain reaction. Therefore, we have developed an oligonucleotide-based microarray using the sequences of 16S–23S rDNA internal transcribed spacer regions (ITS) and the gyrase subunit B gene ( gyrB) found in the most prevalent and devastating waterborne pathogenic agents. from GenBank and edited by using the BioEdit program (Hall, 1999) and ForCon (Raes & Van De Peer, 1999). Consumption of water contaminated with infectious agents, toxic chemicals or radiological hazards represents a significant health risk and is strongly associated with mortality. The safety and accessibility of drinking water are major concerns throughout the world.
